Its causative agent is Newcastle disease virus (NDV), also known as avian paramyxovirus type 1. Many countries maintain a stringent vaccination policy against ND, but there are indications that ND outbreaks can still occur despite intensive vaccination. It has been argued that this may be due to antigenic divergence between the vaccine strains and circulating field strains. Here we present the complete genome sequence of a highly virulent genotype VII virus (NL/93) obtained from vaccinated poultry during an outbreak of ND in the Netherlands in 1992 – 1993. Using this strain, we investigated whether the identified genetic evolution of NDV is accompanied by antigenic evolution. In this study we show that a live vaccine that is antigenically adapted to match the genotype VII NL/93 outbreak strain does not provide increased protection compared to a classic genotype II live vaccine. When challenged with the NL/93 strain, chickens vaccinated with a classic vaccine were completely protected against clinical disease and mortality and virus shedding was significantly reduced, even with a supposedly suboptimal vaccine dose. These results suggest that it is not antigenic variation but rather poor flock immunity due to inadequate vaccination practices that may be responsible for outbreaks and spreading of virulent NDV field strains.
...
-->Introduction
b. RNA isolation, RT-PCR and sequencing
d. Pathogenicity test, haemagglutination inhibition (HI) assay
e. Animals
-->Results
-->Discussion
-->References
...
NEWCASTLE DISEASE VIRUS OUTBREAKS
- VACCINE MISMATCH OR INADEQUATE APPLICATION? -
By Jos C.F.M. Dortmans, Ben P.H. Peeters, Guus Koch, Central Veterinary Institute of Wageningen UR, PO Box 65, 8200 AB Lelystad, The Netherlands
INTRODUCTION
Newcastle disease (ND) is an important infectious disease of poultry because of its potential to cause devastating losses in the poultry industry. Its causative agent is Newcastle disease virus (NDV), also known as avian paramyxovirus type 1, a virus that is able to infect over 240 species of birds and which spreads easily via different routes (Kaleta and Baldauf, 1988). Because ND can cause severe economic losses, it is a notifiable disease to the World Organization for Animal Health (OIE) (Communities, 1992). To keep ND under control, prophylactic vaccination is applied on a large scale in most member states of the European Union and elsewhere in the world.
Most outbreaks of NDV arise in non-vaccinated susceptible animals, and therefore nowadays countries worldwide maintain a stringent vaccination policy. Current NDV vaccines have been used for more than 50 years with proven reputation of safety and efficacy. These vaccine strains belong phylogenetically to the same genotypes (I and II) as viruses isolated in the 1940s, but are divergent from strains that caused ND outbreaks in the last two decades. The most predominant viruses isolated from recent outbreaks in the United States are genotype V viruses (Pedersen et al., 2004), whereas in Africa, Asia and Europe they are mainly of genotype VI (pigeon paramyxovirus type I) and VII (Abolnik et al., 2008; Irvine et al., 2009; Lien et al., 2007; Liu et al., 2007; Yu et al., 2001).
Whilst the available vaccines induce protection against morbidity and mortality from a challenge with highly virulent (velogenic) NDV strains, several studies have shown that they do not prevent infection and virus shedding, which may result in transmission (the infection of other susceptible birds) (Kapczynski and King, 2005; Miller et al., 2009). Although poor vaccine application or quality of the vaccines used cannot be excluded, there are indications that some ND outbreaks occurred despite intensive vaccination against NDV (Abolnik et al., 2004; Bogoyavlenskiy et al., 2009; Hassan et al., 2010; Ke et al., 2010; Oncel et al., 1997; Yang et al., 1999). This might be due to antigenic divergence between the vaccine strain and the circulating field strains (Hu et al., 2009; Kapczynski and King, 2005; Miller et al., 2009, 2007; Qin et al., 2008; van Boven et al., 2008).
The objective of this study was to investigate if the genetic evolution of NDV is accompanied by antigenic evolution which may reduce the efficacy of current ND vaccines. To test this hypothesis, the prophylactic properties of a classic vaccine and a vaccine strain that was adapted to match the genotype of the challenge strain were compared.
^ Top page
.
MATERIALS AND METHODS
a. Cells and viruses
QM5 cells (Antin and Ordahl, 1991) were grown in Ford Dodge QT35 medium (Invitrogen, Carlsbad, CA, USA) at 37oC in a 5% CO2 incubator. Medium was supplemented with 5% fetal bovine serum and 1% of an antibiotic stock consisting of penicillin (100 units/ml) and streptomycin (100 mg/ml).
Strain APMV-1/chicken/NL/152608/93 (NL/93) was isolated during the ND outbreak in the Netherlands in 1992–1993. At that time the intracerebral pathogenicity index (ICPI) was determined to be 1.84. Restriction enzyme and phylogenetic analysis has placed this virus in genotype VII (Lomniczi et al., 1998) or lineage 5a (Aldous et al., 2003). This virus was passaged twice by inoculating 10-day-old embryonated specific-pathogen-free (SPF) eggs to make a new working stock.
The cDNA clone of the low-virulent NDV strain La Sota, named pNDFL and the use of the expression plasmids pCIneo-P and pCIneo-L have been described before (Peeters et al., 1999; Dortmans et al., 2009).
The fowlpox recombinant virus fpEFLT7pol (hereafter called FPV-T7), which expresses the bacteriophage T7 RNA polymerase, was used as previously described (Peeters et al., 1999).
^ Top page
.
b. RNA isolation, RT-PCR and sequencing
Genomic RNA was isolated with a High Pure Viral RNA kit (Roche, Almere, The Netherlands). First-strand DNA synthesis was carried out using a SuperscriptTM III Reverse Transcriptase kit (Invitrogen, Carlsbad, CA, USA). Overlapping sub-genomic cDNA fragments were generated, purified using a High Pure PCR purification kit (Roche, Almere, The Netherlands) and subsequently sequenced.
Primer sequences used for the generation of the overlapping sub-genomic cDNA fragments and for genome sequencing are available upon request. Nucleotide sequencing was carried out using a BigDye Terminator v1.1 cycle sequencing kit and a 3130 genetic analyzer (Applied Biosystems, Nieuwerkerk a/d IJssel, The Netherlands).
^ Top page
.
c. Construction of vaccines
PCR mutagenesis was used to introduce unique restriction sites AscI (position 4527), FseI (position 6347) and PacI (position 8366) into the pNDFL cDNA, resulting in a plasmid designated pNDFLAFP. The F and HN genes of strain NL/93 were synthesized by the GenScript corporation (Piscataway, NJ, USA). The F gene was flanked by the restriction sites AscI and FseI sites and the sequence encoding the protease cleavage site of the F0 protein, 112RRQKR↓F117, was modified into 112GRQGR↓L117 by mutating 5 nucleotides. The HN gene was flanked by the restriction sites FseI and PacI sites. Both genes were cloned between the AscI and PacI sites resulting in the plasmid designated pNDFL(FHN)NL93.
To generate recombinant virus from pNDFLAFP and pNDFL(FHN)NL93, QM5 cells were infected with FPV-T7 and co-transfected with full length genome constructs and plasmids expressing P and L using Fugene HD according to the instructions from the manufacturer (Roche, Mannheim, Germany). After three days, the culture supernatant was harvested, passed through a 0.2 mm filter and subsequently inoculated into 9–11-day-old embryonated specific-pathogen-free (SPF) eggs resulting in viruses NDFLAFP and NDFL(FHN)NL93. To make a virus stock, these viruses were passed through a 0.2 mm filter again and inoculated into 9–11-day old embryonated SPF eggs. Virus production was confirmed by a standard haemagglutination assay.
To verify if the protease cleavage site 112GRQGR↓L117 was not changed back into a velogenic cleavage site, NDFL(FHN)NL93 was passaged 15 times in 9–11-day old embryonated SPF eggs. Viruses from several passages were checked for the ability to form plaques in QM5 cells without the addition of exogenous trypsin and virus of passage 15 was examined by RT-PCR and sequence analysis.
^ Top page
.
d. Pathogenicity test, haemagglutination inhibition (HI) assay
The determination of the intracerebral pathogenicity index in one-day-old chickens and the HI assay were performed as described in the European Community Council Directive 92/66/EEC (Communities, 1992).
^ Top page
.
A total of 45 SPF chickens were bred at the Central Veterinary Institute of Wageningen UR (CVI) and housed in a high containment unit. The experiment was approved by the Ethics Committee for Animal Experiments of the Central Veterinary Institute of Wageningen UR and was in compliance with the Dutch law on animal experiments.
^ Top page
.
f. Experimental design
At three weeks of age four groups of ten SPF chickens were vaccinated by the combined intranasal and intratracheal route. Using a total dose of 106 median egg infectious dose (EID50) per bird, group 1 was vaccinated with NDFL (FHN)NL93 and group 2 with NDFLAFP. Groups 3 and 4 were vaccinated using a total dose of 104 EID50 per bird with NDFL (FHN)NL93 and NDFLAFP, respectively. The negative control group consisted of 5 chickens which received PBS via the same route.
All birds were challenged with strain NL/93 with a total dose of 106 EID50 per chicken two weeks post vaccination. The chickens were observed for clinical signs until the end of the experiment, 14 days post challenge (dpc). Tracheal and cloacal swabs were taken at day 2, 4, 7 and 10 post challenge. The swabs were placed in 2.95% tryptose phosphate buffer with appropriate antibiotics and infectious virus was quantified by serial end-point dilution in 96-well plates using QM5 cells (Antin and Ordahl, 1991).
After three days of incubation at 37oC, cells were fixed and immunological staining (IPMA) was performed to detect the infected cells. For detection, an F protein-specific mouse monoclonal antibody fusie 133 8E12A8C3 (CVI of Wageningen UR) was used as the primary antibody. Rabbit anti-mouse IgG antibodies (DAKO, Heverlee, Belgium) were used as the secondary antibody. Activity of peroxidase was detected using 3-amino-9-ethyl-carbazole (Sigma, St. Louis, MO, USA) as the substrate. Geometric mean virus titres were calculated using the method of Reed and Muench (Reed and Muench, 1938) and expressed as log10TCID50/ml. QM5-negative samples were tested for the presence of virus by inoculation of undiluted tissue homogenates in embryonated SPF eggs.
Serum samples were obtained at day ‒ 14, day 0 and day 14 after challenge and the HI assay was performed using 8 haemagglutinating units of antigen. The geometric mean antibody titres were expressed on a log2 scale. The antigens used in this experiment were NDFLAFP, NDFL (FHN)NL93, Netherlands/93 and strain Ulster. The latter is used as a standard antigen in the HI test that is routinely used in ND diagnostic laboratories.
^ Top page
.
For differences in frequencies of positive trachea and cloacal swabs, morbidity and mortality of the vaccinated group versus the non-vaccinated group Fisher’s exact test was used. Statistical analysis of virus titres and serology titres were employed using the Wilcoxon Mann–Whitney Test. Differences were considered to be significant when P < 0.05.
^ Top page
.
RESULTS
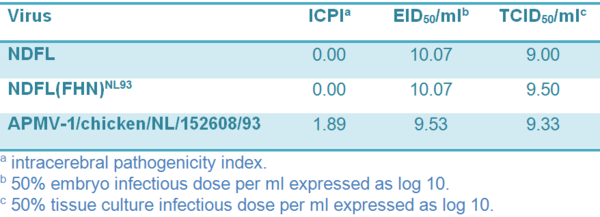
The complete genome sequence of APMV-1/chicken/ NL/152608/93 (NL/93) was determined. The assembled sequence consists of 15,192 nt (GenBank accession number JN986837), which puts it into class II according to the classification based on genome size. Comparison of the genome sequence of NL/93 with those of APMV-1. strains available in GenBank showed similarities with ND viruses isolated from ducks and geese. The highest sequence identity (95%) was found with strain ND/03/044 (accession number GQ338310) that was originally isolated in China. Phylogenetic analysis confirmed previous studies which showed that NL/93 can be classified as genotype VIIa (Lomniczi et al., 1998; Aldous et al., 2003). Grouping was based on the partial (nt 47–422) fusion protein sequence and the complete genome sequence (data not shown). The pathogenicity index amounted to 1.89 (Table 1).
Table 1 Virus properties
In order to generate a new NDV vaccine that is genetically more similar to current genotype VII outbreak viruses, we used the backbone of the La Sota strain, NDFL, to generate a virus containing the F and HN genes of NL/93. Because NL/93 contains a velogenic cleavage site motif in its F protein, we changed it into a lentogenic cleavage site as described in Section 2. The resulting virus NDFL(FHN)NL93 could be rescued and sequence analysis showed that the virus did not contain any unintended mutations. The ICPI was determined to be 0.0.
To evaluate the genetic stability of the modified cleavage site of the F gene, the recombinant virus was serially passaged 15 times in 9–11-day old embryonated SPF eggs. Virus of passage 15 was not able to form plaques in QM5 cells without the addition of exogenous trypsin (indicative of a lentogenic phenotype) and sequence analysis showed that the F and HN genes remained unaltered (data not shown).
At the age of 18–21 days, groups of 10 SPF chickens were vaccinated intranasally / intratracheally with a dose of 106 EID50 (optimal dose) or 104 EID50 (suboptimal) NDFL(FHN)NL93 or NDFLAFP. As a control, 5 SPF chickens were inoculated with PBS. Fourteen days post vaccination (0 dpc) all birds were challenged with 106 EID50 NL/93.
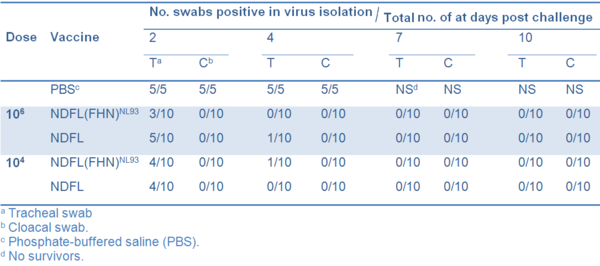
No clinical symptoms were observed in any of the vaccinated chickens and none died. In contrast all birds in the non-vaccinated control group died between 4 and 7 dpc after developing severe clinical signs. All nonvaccinated chickens shed virus from both trachea and cloaca, whereas none of the vaccinated birds shed detectable virus from cloaca. Only a few birds in each vaccinated group shed virus from trachea on 2 and 4 dpc (Table 2).
Table 2 Frequency of isolation of challenge virus in different vaccine groups
The geometric mean virus titre of the challenge control group was 104.7 at 2 dpc and 106.1 at 4 dpc in trachea swabs and 101.0 at 2 dpc and 103.9 at 4 dpc in cloaca swabs (Figure 1). Animals in the vaccinated groups (taking into account only birds that shed virus, see Table 2) shed significantly lower amounts of virus compared to the challenge control group (P < 0.05), whereas the differences between the different vaccinated groups were not significant. These results were confirmed by estimating and adding up the area under the virus excretion curve of each individual animal per group (data not shown).
Figure 1 Geometric mean virus titres determined in tracheal and cloacal swabs at 2 and 4 dpc of chickens that had detectable virus titres. Small letters (a–e) represent the amount of viral shedding chickens in the total group (see also Table 2):
- (a) 5 out of the 5
- (b) 3 out of 10
- (c) 5 out of 10
- (d) 4 out of 10
- (e) 1 out of 10
Titres (log10TCID50/ml) were determined by serial end-point dilution in 96-well plates using QM5 cells. Error bars represent standard deviations. Viral shedding in all vaccinated groups is significantly reduced compared to the PBS inoculated group (P < 0.05).
Antibody titres were determined in the HI test using the following viruses as antigen: Ulster, NL/93, NDFLAFP and NDFL(FHN)NL93. The amino acid similarities of the F and HN proteins of the four viruses are shown in Table 3.
Table 3 Similarity between strain used in this studya
The results of the HI test (Figure 2) showed that there were no significant differences in geometric mean antibody titres between the two vaccines and between the high and low dose groups at either time point when each data pair from the different antigens was compared. Generally, no differences in antibody titres could be observed when the different antigens were compared. However, a statistically significantly difference (P < 0.05) in mean HI titre was detected in the group of chickens vaccinated with 106 NDFL(FHN)NL93 between the antigens Ulster and NL/93 at 0 dpc, and in the group of chickens vaccinated with 104 NDFL(FHN)NL93, between the antigens Ulster and NDFL(FHN)NL93 (Figure 2). Seroconversion, defined as an HI titre increase of at least 2 log2 after challenge was observed in 6 of 10 chickens vaccinated with 106 NDFL(FHN)NL93 and 106 NDFLAFP, 10 of 10 in chickens vaccinated with 104 NDFL(FHN)NL93, and 4 of 10 chickens vaccinated with 104 NDFLAFP.
Figure 2 Serology of SPF chickens vaccinated with the two vaccines with doses 106 and 104 EID50. Geometric mean HI titres (log2) were determined at 0 and 14 dpc using four different antigens. Error bars represent standard deviations. An asterisk (*) represents a significant difference in antibody titre using different antigens (P < 0.05). The legend shows the antigens used in the HI test.
In summary, it can be concluded that the HI titre after experimental vaccination with both a homologous and a heterologous vaccine is very high, even with a vaccine dose that is 500 – 5.000 times lower than what is considered optimal (Allan et al., 1978). All vaccinated chickens were protected against mortality and disease, and viral shedding from the trachea was eliminated in the majority of birds or significantly reduced in the remainder. None of the vaccinated birds shed virus from the cloaca.
^ Top page
.
DISCUSSION
In the present study, we have shown that a live vaccine that is adapted to an outbreak strain does not increase protection nor reduce viral shedding compared to a classic live vaccine. In fact the classic live vaccine, based on the lentogenic La Sota strain, performs very well even with a supposedly suboptimal vaccination dose of 104 EID50.
When challenged with a velogenic virus, i.e. the outbreak strain NL/93, chickens are completely protected against clinical disease and mortality and viral shedding is absent or significantly reduced. The adapted vaccine, containing the envelope glycoproteins F and HN of genotype VII virus NL/93, showed the same results compared to the La Sota based vaccine. These results are in line with several other studies (Bwala et al., 2009; Jeon et al., 2008; Liu et al., 2003). Cross protection studies showed a 100% protection efficacy when vaccinated (genotype II) chickens were challenged with heterologous viruses of genotypes V, VIb, VIg, VIId and IX (Bwala et al., 2009; Liu et al., 2003).
Unfortunately, both studies did not examine viral shedding after challenge to evaluate the level of protection. Jeon et al. did observe viral shedding by vaccinated chickens after challenge with a genotype VIId virus, but this was significantly reduced when compared to unvaccinated chickens (Jeon et al., 2008). The authors concluded that chickens with lower vaccine-induced immunity may continue to be susceptible to NDV infection. This may explain why field viruses are able to circulate within farms in which chickens are vaccinated suboptimally.
Our results do not support the concerns expressed in several other published reports (Hu et al., 2009; Kapczynski and King, 2005; Liang et al., 2002; Miller et al., 2007, 2009; van Boven et al., 2008; Yu et al., 2001) which showed that current vaccines prevent disease but cannot stop viral shedding. When genotype-matched vaccines were used, viral shedding was significantly reduced compared to genotype-mismatched vaccines (Hu et al., 2009; Miller et al., 2009). The question is whether these findings have a real impact on future outbreaks, since no transmission experiments have been performed to test these viral shedding differences with genotype-matched and genotype- mismatched vaccines.
It has been proposed that genotype VII viruses might have originated in the Far East and that the earliest isolation was in Taiwan in 1984 (Lomniczi et al., 1998; Yang et al., 1999). In the early nineties, viruses isolated from outbreaks in Europe belong to genotype VII (Lomniczi et al., 1998). One of these viruses is the Dutch strain NL/93 from the outbreak in 1992–1993. However, similar viruses are still a threat in the poultry industry worldwide.
Genotype VII viruses have been isolated in the European union in the last decade (Alexander, 2011), Kazakhstan and Kyrgyzstan (Bogoyavlenskiy et al., 2009), Sudan (Hassan et al., 2010), Malaysia (Tan et al., 2010), South Korea (Lee et al., 2004), Taiwan (Ke et al., 2010; Lien et al., 2007), China (Liu et al., 2007), South Africa (Abolnik et al., 2004) and West Africa (Snoeck et al., 2009).
In the field, routine vaccinations do not result in sufficient immunity to prevent transmission (van Boven et al., 2008). Poor vaccination practices and / or concurrent infection with immunosuppressive pathogens rather than antigenic variation may be responsible for poor immunity levels giving the way for viral spreading (Jeon et al., 2008).
Moreover, ND vaccination often causes minimal to sever side effects causing production losses and vaccination practices are therefore directed at avoiding these reactions rather than at obtaining optimal immunity. Alternatively, it may be that the currently practiced immunization methods involving vaccine application by spraying (aerosol) or the addition to drinking water are not robust enough to guarantee that each birds receives an adequate vaccine dose. If a new vaccine is still handled poorly or vaccination schedules are inappropriate, a genotype-matched vaccine still will not give optimal protection.
As reported before, more studies will have to be performed to test the above-mentioned statements.
^ Top page
.
...
REFERENCES
Abolnik, C., Gerdes, G.H., Kitching, J., Swanepoel, S., Romito, M., Bisschop, S.P., 2008. Characterization of pigeon paramyxoviruses (Newcastle disease virus) isolated in South Africa from 2001 to 2006. Onderstepoort J. Vet. 75, 147–152.
Abolnik, C., Horner, R.F., Bisschop, S.P., Parker, M.E., Romito, M., Viljoen, G.J., 2004. A phylogenetic study of South African Newcastle disease virus strains isolated between 1990 and 2002 suggests epidemiological origins in the Far East. Arch. Virol. 149, 603–619.
Aldous, E.W., Mynn, J.K., Banks, J., Alexander, D.J., 2003. A molecular epidemiological study of avian paramyxovirus type 1 (Newcastle disease virus) isolates by phylogenetic analysis of a partial nucleotide sequence of the fusion protein gene. Avian Pathol. 32, 239–256.
Alexander, D.J., 2011. Newcastle disease in the European Union 2000 to 2009. Avian Pathol. 40, 547–558.
Allan, W.H., Lancaster, J.E., Toth, B., 1978. Newcastle disease vaccines. Their production and use. Food and Agriculture Organization of the United Nations, Rome, 163 pp.
Antin, P.B., Ordahl, C.P., 1991. Isolation and characterization of an avian myogenic cell line. Dev. Biol. 143, 111–121.
Bogoyavlenskiy, A., Berezin, V., Prilipov, A., Usachev, E., Lyapina, O., Korotetskiy, I., Zaitceva, I., Asanova, S., Kydyrmanov, A., Daulbaeva, K., Shakhvorostova, L., Sayatov, M., King, D., 2009. Newcastle disease outbreaks in Kazakhstan and Kyrgyzstan during 1998, 2000, 2001, 2003, 2004, and 2005 were caused by viruses of the genotypes VIIb and VIId. Virus Genes 39, 94–101.
Bwala, D.G., Abolnik, C., van Wyk, A., Cornelius, E., Bisschop, S.P., 2009. Efficacy of a genotype 2 Newcastle disease vaccine (Avinew) against challenge with highly virulent genotypes 5d and 3d. J. S. Afr. Vet. Assoc. 80, 174–178.
Communities, C.o.t.E., 1992. Council directive 92/66/EEC of 14 July 1992 introducing community measures for the control of Newcastle disease. Offic. J. Eur. Communities L260, 1–20.
Dortmans, J.C.F.M., Koch, G., Rottier, P.J.M., Peeters, B.P.H., 2009. Virulence of pigeon paramyxovirus type 1 (PPMV-1) does not always correlate with the cleavability of its fusion protein. J. Gen. Virol. 90, 2746–2750.
Hassan, W., Khair, S.A., Mochotlhoane, B., Abolnik, C., 2010. Newcastle disease outbreaks in the Sudan from 2003 to 2006 were caused by viruses of genotype 5d. Virus Genes 40, 106–110.
Hu, S., Ma, H., Wu, Y., Liu, W., Wang, X., Liu, Y., Liu, X., 2009. A vaccine candidate of attenuated genotype VII Newcastle disease virus generated by reverse genetics. Vaccine 27, 904–910.
Irvine, R.M., Aldous, E.W., Manvell, R.J., Cox, W.J., Ceeraz, V., Fuller, C.M., Wood, A.M., Milne, J.C., Wilson, M., Hepple, R.G., Hurst, A., Sharpe, C.E., Alexander, D.J., Brown, I.H., 2009. Outbreak of Newcastle disease due to pigeon paramyxovirus type 1 in grey partridges (Perdix perdix) in Scotland in October 2006. Vet. Rec. 165, 531–535.
Jeon, W.J., Lee, E.K., Lee, Y.J., Jeong, O.M., Kim, Y.J., Kwon, J.H., Choi, K.S., 2008. Protective efficacy of commercial inactivated Newcastle disease virus vaccines in chickens against a recent Korean epizootic strain. J. Vet. Sci. 9, 295–300.
Kaleta, E.F., Baldauf, C., 1988. Newcastle disease in free-living and pet birds. In: Alexander, D.J. (Ed.), Newcastle Disease. Kluwer Academic Publishers, Boston, pp. 197–246.
Kapczynski, D.R., King, D.J., 2005. Protection of chickens against overt clinical disease and determination of viral shedding following vaccination with commercially available Newcastle disease virus vaccines upon challenge with highly virulent virus from the California 2002 exotic Newcastle disease outbreak. Vaccine 23, 3424–3433.
Ke, G.M., Yu, S.W., Ho, C.H., Chu, P.Y., Ke, L.Y., Lin, K.H., Tsai, Y.C., Liu, H.J., Lin, M.Y., 2010. Characterization of newly emerging Newcastle disease viruses isolated during 2002–2008 in Taiwan. Virus Res. 147, 247–257.
Lee, Y.J., Sung, H.W., Choi, J.G., Kim, J.H., Song, C.S., 2004. Molecular epidemiology of Newcastle disease viruses isolated in South Korea using sequencing of the fusion protein cleavage site region and phylogenetic relationships. Avian Pathol. 33, 482–491.
Liang, R., Cao, D.J., Li, J.Q., Chen, J., Guo, X., Zhuang, F.F., Duan, M.X., 2002. Newcastle disease outbreaks in western China were caused by the genotypes VIIa and VIII. Vet. Microbiol. 87, 193–203.
Lien, Y.Y., Lee, J.W., Su, H.Y., Tsai, H.J., Tsai, M.C., Hsieh, C.Y., Tsai, S.S., 2007. Phylogenetic characterization of Newcastle disease viruses isolated in Taiwan during 2003–2006. Vet. Microbiol. 123, 194–202.
Liu, H., Wang, Z., Wu, Y., Zheng, D., Sun, C., Bi, D., Zuo, Y., Xu, T., 2007. Molecular epidemiological analysis of Newcastle disease virus isolated in China in 2005. J. Virol. Methods 140, 206–211.
Liu, X.F., Wan, H.Q., Ni, X.X., Wu, Y.T., Liu, W.B., 2003. Pathotypical and genotypical characterization of strains of Newcastle disease virus isolated from outbreaks in chicken and goose flocks in some regions of China during 1985–2001. Arch. Virol. 148, 1387–1403.
Lomniczi, B., Wehmann, E., Herczeg, J., Ballagi-Pordany, A., Kaleta, E.F., Werner, O., Meulemans, G., Jorgensen, P.H., Mante, A.P., Gielkens, A.L., Capua, I., Damoser, J., 1998. Newcastle disease outbreaks in recent years in western Europe were caused by an old (VI) and a novel genotype (VII). Arch. Virol. 143, 49–64.
Miller, P.J., Estevez, C., Yu, Q., Suarez, D.L., King, D.J., 2009. Comparison of viral shedding following vaccination with inactivated and live Newcastle disease vaccines formulated with wild-type and recombinant viruses. Avian Dis. 53, 39–49.
Miller, P.J., King, D.J., Afonso, C.L., Suarez, D.L., 2007. Antigenic differences among Newcastle disease virus strains of different genotypes used in vaccine formulation affect viral shedding after a virulent challenge. Vaccine 25, 7238–7246.
Oncel, T., Alexander, D.J., Manvell, R.J., Ture, O., 1997. Characterization of Newcastle disease viruses isolated from chickens and pigeons in the South Marmara region of Turkey. Avian Pathol. 26, 129–137.
Pedersen, J.C., Senne, D.A., Woolcock, P.R., Kinde, H., King, D.J., Wise, M.G., Panigrahy, B., Seal, B.S., 2004. Phylogenetic relationships among virulent Newcastle disease virus isolates from the 2002–2003 outbreak in California and other recent outbreaks in North America. J. Clin. Microbiol. 42, 2329–2334.
Peeters, B.P., de Leeuw, O.S., Koch, G., Gielkens, A.L., 1999. Rescue of Newcastle disease virus from cloned cDNA: evidence that cleavability of the fusion protein is a major determinant for virulence. J. Virol. 73, 5001–5009.
Qin, Z.M., Tan, L.T., Xu, H.Y., Ma, B.C., Wang, Y.L., Yuan, X.Y., Liu, W.J., 2008. Pathotypical characterization and molecular epidemiology of Newcastle disease virus isolates from different hosts in China from 1996 to 2005. J. Clin. Microbiol. 46, 601–611.
Reed, L.J., Muench, H.A., 1938. A simple method of estimating fifty percent endpoints. Am. J. Hyg. 27, 493–497.
Snoeck, C.J., Ducatez, M.F., Owoade, A.A., Faleke, O.O., Alkali, B.R., Tahita, M.C., Tarnagda, Z., Ouedraogo, J.B., Maikano, I., Mbah, P.O., Kremer, J.R., Muller, C.P., 2009. Newcastle disease virus in West Africa: new virulent strains identified in non-commercial farms. Arch. Virol. 154, 47–54.
Tan, S.W., Ideris, A., Omar, A.R., Yusoff, K., Hair-Bejo, M., 2010. Sequence and phylogenetic analysis of Newcastle disease virus genotypes isolated in Malaysia between 2004 and 2005. Arch. Virol. 155, 63–70.
van Boven, M., Bouma, A., Fabri, T.H., Katsma, E., Hartog, L., Koch, G., 2008. Herd immunity to Newcastle disease virus in poultry by vaccination. Avian Pathol. 37, 1–5.
Yang, C.Y., Shieh, H.K., Lin, Y.L., Chang, P.C., 1999. Newcastle disease virus isolated from recent outbreaks in Taiwan phylogenetically related to viruses (genotype VII) from recent outbreaks in western Europe. Avian Dis. 43, 125–130.
Yu, L., Wang, Z., Jiang, Y., Chang, L., Kwang, J., 2001. Characterization of newly emerging Newcastle disease virus isolates from the People’s Republic of China and Taiwan. J. Clin. Microbiol. 39, 3512–3519.
...
This article appeared in Veterinary Microbiology 160 (2012) 17–22,Received in revised form 1 May 2012 Accepted 7 May 2012.©Copyright 2012, All Rights Reserved.

 Corporate Website
Corporate Website
 Africa
Africa
 Argentina
Argentina
 Asia
Asia
 Australia
Australia
 Belgium
Belgium
 Brazil
Brazil
 Bulgaria
Bulgaria
 Canada (EN)
Canada (EN)
 Chile
Chile
 China
China
 Colombia
Colombia
 Denmark
Denmark
 Egypt
Egypt
 France
France
 Germany
Germany
 Greece
Greece
 Hungary
Hungary
 Indonesia
Indonesia
 Italia
Italia
 India
India
 Japan
Japan
 Korea
Korea
 Malaysia
Malaysia
 Mexico
Mexico
 Middle East
Middle East
 Netherlands
Netherlands
 Peru
Peru
 Philippines
Philippines
 Poland
Poland
 Portugal
Portugal
 Romania
Romania
 Russia
Russia
 South Africa
South Africa
 Spain
Spain
 Sweden
Sweden
 Thailand
Thailand
 Tunisia
Tunisia
 Turkey
Turkey
 Ukraine
Ukraine
 United Kingdom
United Kingdom
 USA
USA
 Vietnam
Vietnam